Biobank Intro Series: Getting Started
How to do biobank analysis without losing your mind
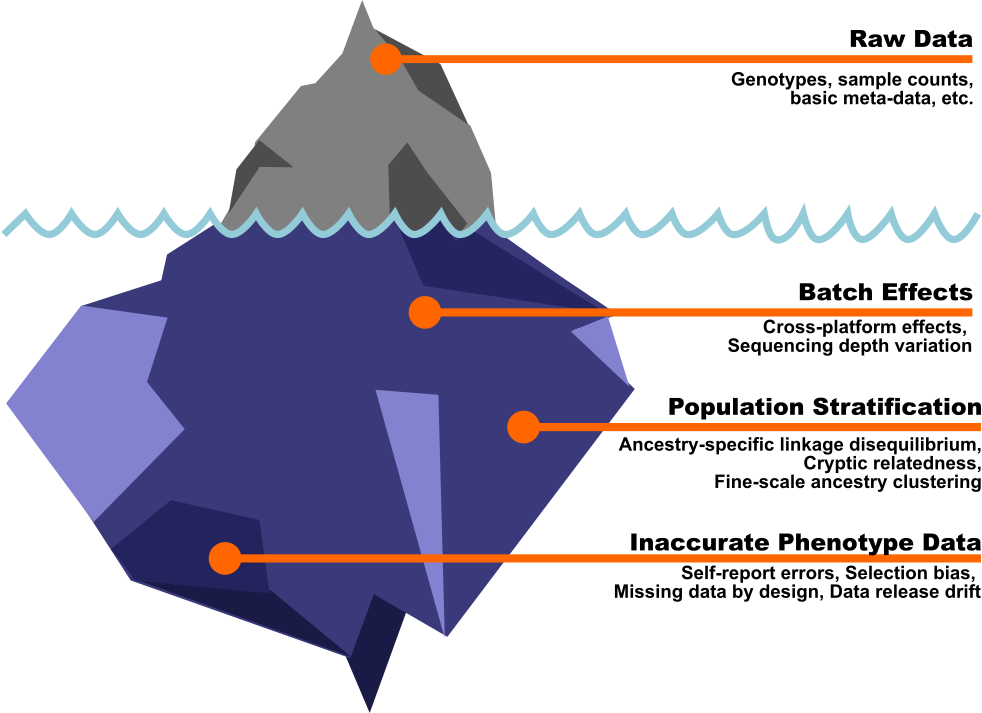
If I learned anything during my first few months as a bioinformatics contractor, it’s that you need to know your question before you worry about the hidden complexities. Like icebergs, biobanks have immense power—and like icebergs, their foundation is made up of massive complexities beneath the surface that can sink your analysis if you’re not prepared. (The Titanic had better odds.)
The first step to any biobank analysis is strategic rather than technical.
- What specific research question are you trying to answer?
- Who is your audience?
That’s it. Two questions. Answer these before you touch a single line of code.
Consider my own work as an example:
During my postdoc: I was investigating the genetic architecture of cardiomyopathy for publication. My goal was to understand how genetic variants contribute to disease risk. A large challenge, working within the context of rare disease, is overcoming limited statistical power. Therefore, to strengthen the association signal, I restricted analyses to individuals of White British ancestry (UK Biobank data field 22006, coding 1). The healthy volunteer recruitment biases and population structure weren’t just footnotes in the discussion section—they fundamentally shaped what I could and couldn’t claim about cardiomyopathy genetics. I needed to understand the people making up the dataset as deeply as I understood the variants themselves.
As a contractor: I provide bioinformatics support for hypothesis-driven study design that could lead to drug development. The client wants to find out if a genetic signal associates with a phenotype strongly enough to justify further investigation. In other words, I’m running pilot analyses. I need clean cohort definitions, appropriate statistical methods, and enough power to detect effects. The recruitment biases? Important to document, but they don’t invalidate finding a biological association in this specific dataset. My job is to answer the client’s question efficiently and move the project forward.
In my contractor role, I’ve learned that curiosity must be disciplined. My job is to hit project checkpoints and answer the client’s specific question, not to chase down every interesting technical rabbit hole. The data complexity matters, but staying on track isn’t just good project management; it’s professional integrity.
Only after you’ve defined your question and audience can you identify which data complexities actually matter. Every dataset comes with biases and baggage, and I fight two competing urges:
-
The paranoid perfectionist: Document every limitation, explore every confounding variable, and fall into a six-month rabbit hole about batch effects
-
The YOLO analyst: Ignore all complexity, run the analysis, get wildly misleading results, and have to redo everything anyway.
Neither instinct serves the project. The discipline I’ve had to learn is this: acknowledge the complexity but focus only on what affects your specific research question.
The good news is that this complexity isn’t insurmountable. It just requires understanding the data first. I’ve already fallen into some of these rabbit holes, so hopefully you won’t have to. In a series of posts, I hope to demystify UK Biobank and All of Us so you can navigate them confidently. But let me be clear, I cannot cover every nuance of these massive resources. Your specific research question may surface complexities I haven’t encountered. What I can give you are foundational issues that matter across most use cases, the patterns to watch for, and questions to ask.