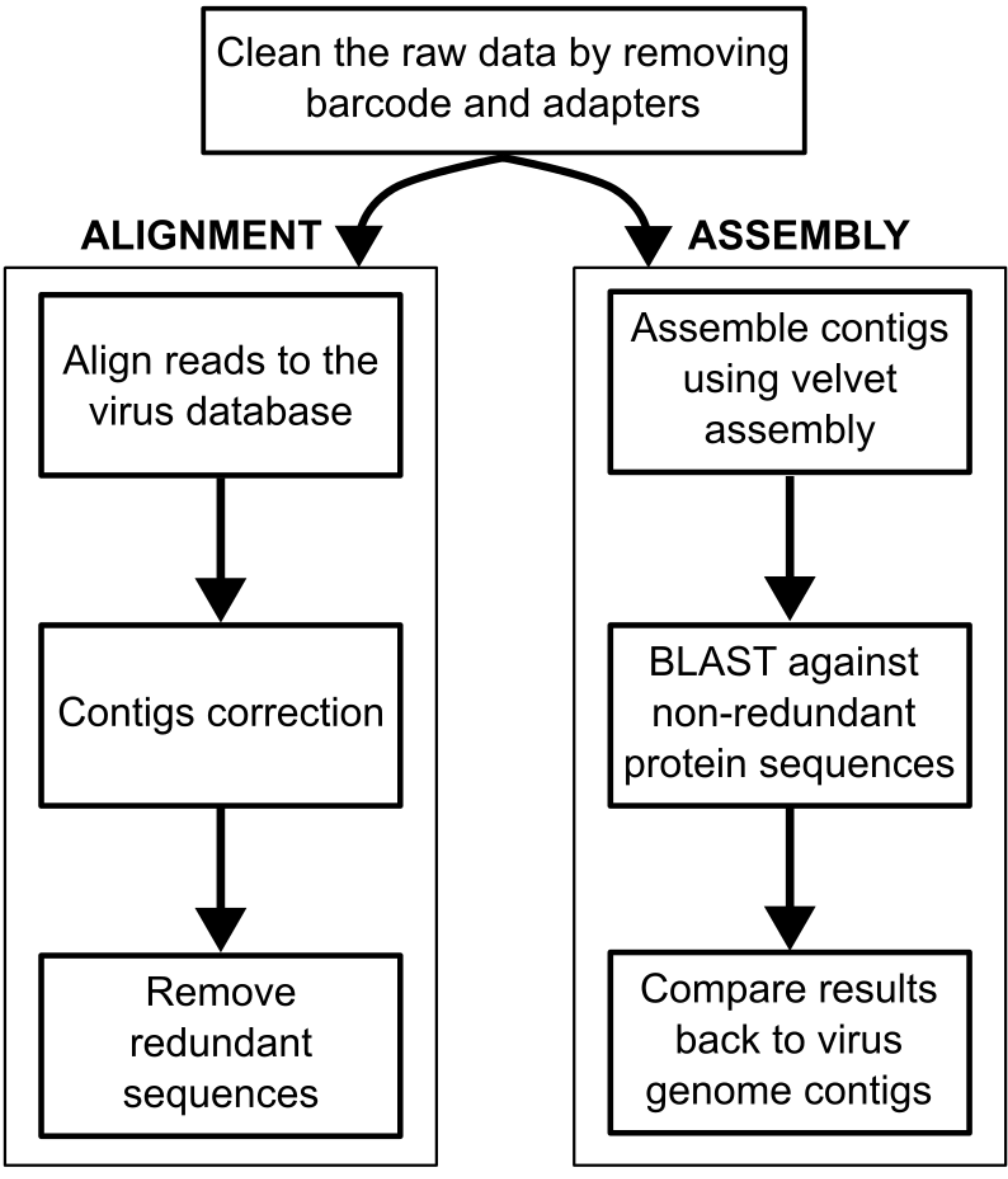
Determining the pan-African sweetpotato virome: Understanding virus diversity, distribution and evolution and their impacts on sweetpotato production in Africa
Gutierrez, D., Tachin, M., Schulz, S., Miano, D., Ndunguru, J., Mukassa, S., ... & Kreuze, J. F.
CIP Research Report • 2012